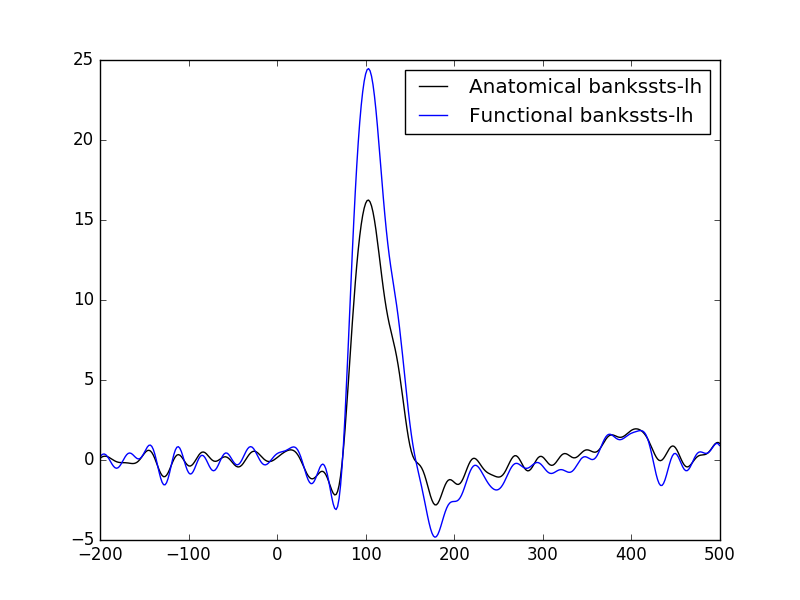
Threshold source estimates and produce a functional label. The label is typically the region of interest that contains high values. Here we compare the average time course in the anatomical label obtained by FreeSurfer segmentation and the average time course from the functional label. As expected the time course in the functional label yields higher values.
# Author: Luke Bloy <luke.bloy@gmail.com>
# Alex Gramfort <alexandre.gramfort@telecom-paristech.fr>
# License: BSD (3-clause)
import numpy as np
import matplotlib.pyplot as plt
import mne
from mne.minimum_norm import read_inverse_operator, apply_inverse
from mne.datasets import sample
print(__doc__)
data_path = sample.data_path()
subjects_dir = data_path + '/subjects'
fname_inv = data_path + '/MEG/sample/sample_audvis-meg-oct-6-meg-inv.fif'
fname_evoked = data_path + '/MEG/sample/sample_audvis-ave.fif'
subjects_dir = data_path + '/subjects'
subject = 'sample'
snr = 3.0
lambda2 = 1.0 / snr ** 2
method = "dSPM" # use dSPM method (could also be MNE or sLORETA)
# Compute a label/ROI based on the peak power between 80 and 120 ms.
# The label bankssts-lh is used for the comparison.
aparc_label_name = 'bankssts-lh'
tmin, tmax = 0.080, 0.120
# Load data
evoked = mne.read_evokeds(fname_evoked, condition=0, baseline=(None, 0))
inverse_operator = read_inverse_operator(fname_inv)
src = inverse_operator['src'] # get the source space
# Compute inverse solution
stc = apply_inverse(evoked, inverse_operator, lambda2, method,
pick_ori='normal')
# Make an STC in the time interval of interest and take the mean
stc_mean = stc.copy().crop(tmin, tmax).mean()
# use the stc_mean to generate a functional label
# region growing is halted at 60% of the peak value within the
# anatomical label / ROI specified by aparc_label_name
label = mne.read_labels_from_annot(subject, parc='aparc',
subjects_dir=subjects_dir,
regexp=aparc_label_name)[0]
stc_mean_label = stc_mean.in_label(label)
data = np.abs(stc_mean_label.data)
stc_mean_label.data[data < 0.6 * np.max(data)] = 0.
func_labels, _ = mne.stc_to_label(stc_mean_label, src=src, smooth=True,
subjects_dir=subjects_dir, connected=True)
# take first as func_labels are ordered based on maximum values in stc
func_label = func_labels[0]
# load the anatomical ROI for comparison
anat_label = mne.read_labels_from_annot(subject, parc='aparc',
subjects_dir=subjects_dir,
regexp=aparc_label_name)[0]
# extract the anatomical time course for each label
stc_anat_label = stc.in_label(anat_label)
pca_anat = stc.extract_label_time_course(anat_label, src, mode='pca_flip')[0]
stc_func_label = stc.in_label(func_label)
pca_func = stc.extract_label_time_course(func_label, src, mode='pca_flip')[0]
# flip the pca so that the max power between tmin and tmax is positive
pca_anat *= np.sign(pca_anat[np.argmax(np.abs(pca_anat))])
pca_func *= np.sign(pca_func[np.argmax(np.abs(pca_anat))])
Out:
Reading /home/ubuntu/mne_data/MNE-sample-data/MEG/sample/sample_audvis-ave.fif ...
Read a total of 4 projection items:
PCA-v1 (1 x 102) active
PCA-v2 (1 x 102) active
PCA-v3 (1 x 102) active
Average EEG reference (1 x 60) active
Found the data of interest:
t = -199.80 ... 499.49 ms (Left Auditory)
0 CTF compensation matrices available
nave = 55 - aspect type = 100
Projections have already been applied. Setting proj attribute to True.
Applying baseline correction (mode: mean)
Reading inverse operator decomposition from /home/ubuntu/mne_data/MNE-sample-data/MEG/sample/sample_audvis-meg-oct-6-meg-inv.fif...
Reading inverse operator info...
[done]
Reading inverse operator decomposition...
[done]
305 x 305 full covariance (kind = 1) found.
Read a total of 4 projection items:
PCA-v1 (1 x 102) active
PCA-v2 (1 x 102) active
PCA-v3 (1 x 102) active
Average EEG reference (1 x 60) active
Noise covariance matrix read.
22494 x 22494 diagonal covariance (kind = 2) found.
Source covariance matrix read.
22494 x 22494 diagonal covariance (kind = 6) found.
Orientation priors read.
22494 x 22494 diagonal covariance (kind = 5) found.
Depth priors read.
Did not find the desired covariance matrix (kind = 3)
Reading a source space...
Computing patch statistics...
Patch information added...
Distance information added...
[done]
Reading a source space...
Computing patch statistics...
Patch information added...
Distance information added...
[done]
2 source spaces read
Read a total of 4 projection items:
PCA-v1 (1 x 102) active
PCA-v2 (1 x 102) active
PCA-v3 (1 x 102) active
Average EEG reference (1 x 60) active
Source spaces transformed to the inverse solution coordinate frame
Preparing the inverse operator for use...
Scaled noise and source covariance from nave = 1 to nave = 55
Created the regularized inverter
Created an SSP operator (subspace dimension = 3)
Created the whitener using a full noise covariance matrix (3 small eigenvalues omitted)
Computing noise-normalization factors (dSPM)...
[done]
Picked 305 channels from the data
Computing inverse...
(eigenleads need to be weighted)...
(dSPM)...
[done]
Reading labels from parcellation..
read 1 labels from /home/ubuntu/mne_data/MNE-sample-data/subjects/sample/label/lh.aparc.annot
read 0 labels from /home/ubuntu/mne_data/MNE-sample-data/subjects/sample/label/rh.aparc.annot
[done]
-- number of connected vertices : 8196
8.5% of original source space vertices have been omitted, tri-based connectivity will have holes.
Consider using distance-based connectivity or morphing data to all source space vertices.
Reading labels from parcellation..
read 1 labels from /home/ubuntu/mne_data/MNE-sample-data/subjects/sample/label/lh.aparc.annot
read 0 labels from /home/ubuntu/mne_data/MNE-sample-data/subjects/sample/label/rh.aparc.annot
[done]
Extracting time courses for 1 labels (mode: pca_flip)
Extracting time courses for 1 labels (mode: pca_flip)
plot the time courses….
plt.figure()
plt.plot(1e3 * stc_anat_label.times, pca_anat, 'k',
label='Anatomical %s' % aparc_label_name)
plt.plot(1e3 * stc_func_label.times, pca_func, 'b',
label='Functional %s' % aparc_label_name)
plt.legend()
plt.show()
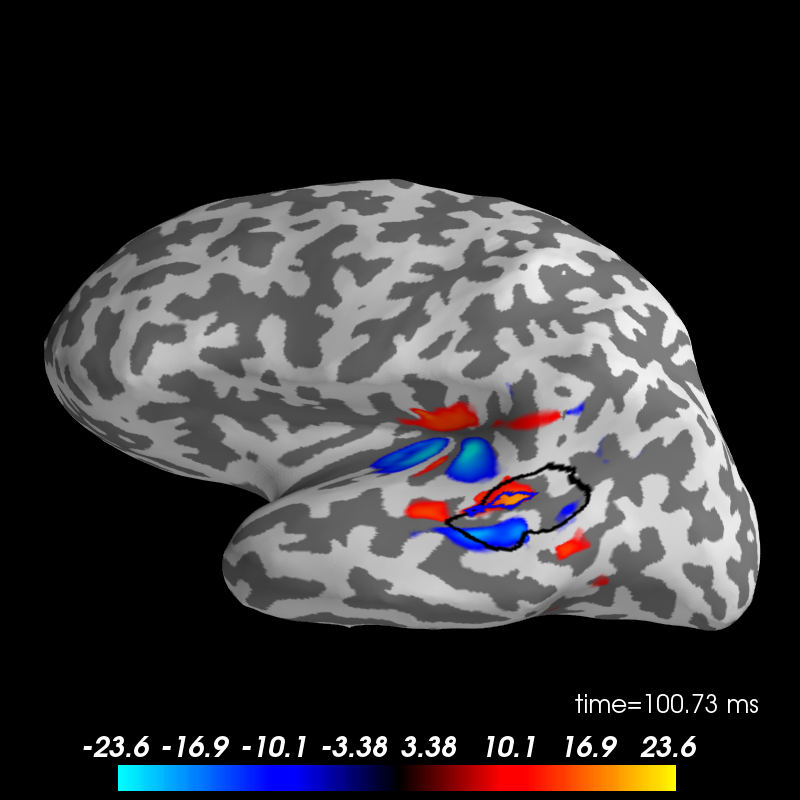
plot brain in 3D with PySurfer if available
brain = stc_mean.plot(hemi='lh', subjects_dir=subjects_dir)
brain.show_view('lateral')
# show both labels
brain.add_label(anat_label, borders=True, color='k')
brain.add_label(func_label, borders=True, color='b')
Out:
Updating smoothing matrix, be patient..
Smoothing matrix creation, step 1
Smoothing matrix creation, step 2
Smoothing matrix creation, step 3
Smoothing matrix creation, step 4
Smoothing matrix creation, step 5
Smoothing matrix creation, step 6
Smoothing matrix creation, step 7
Smoothing matrix creation, step 8
Smoothing matrix creation, step 9
Smoothing matrix creation, step 10
colormap: fmin=-2.36e+01 fmid=0.00e+00 fmax=2.36e+01 transparent=0
Total running time of the script: ( 0 minutes 21.148 seconds)