Here we compute the evoked from raw for the Brainstorm CTF phantom tutorial dataset. For comparison, see [1] and:
| [1] | Tadel F, Baillet S, Mosher JC, Pantazis D, Leahy RM. Brainstorm: A User-Friendly Application for MEG/EEG Analysis. Computational Intelligence and Neuroscience, vol. 2011, Article ID 879716, 13 pages, 2011. doi:10.1155/2011/879716 |
# Authors: Eric Larson <larson.eric.d@gmail.com>
#
# License: BSD (3-clause)
import os.path as op
import numpy as np
import matplotlib.pyplot as plt
import mne
from mne import fit_dipole
from mne.datasets.brainstorm import bst_phantom_ctf
from mne.io import read_raw_ctf
print(__doc__)
The data were collected with a CTF system at 2400 Hz.
data_path = bst_phantom_ctf.data_path()
# Switch to these to use the higher-SNR data:
# raw_path = op.join(data_path, 'phantom_200uA_20150709_01.ds')
# dip_freq = 7.
raw_path = op.join(data_path, 'phantom_20uA_20150603_03.ds')
dip_freq = 23.
erm_path = op.join(data_path, 'emptyroom_20150709_01.ds')
raw = read_raw_ctf(raw_path, preload=True)
Out:
ds directory : /home/ubuntu/mne_data/MNE-brainstorm-data/bst_phantom_ctf/phantom_20uA_20150603_03.ds
res4 data read.
hc data read.
Separate EEG position data file read.
Quaternion matching (desired vs. transformed):
-0.39 74.35 0.00 mm <-> -0.39 74.35 0.00 mm (orig : -39.23 63.82 -204.07 mm) diff = 0.000 mm
0.39 -74.35 0.00 mm <-> 0.39 -74.35 0.00 mm (orig : 65.69 -41.53 -205.68 mm) diff = 0.000 mm
75.00 0.00 0.00 mm <-> 75.00 -0.00 0.00 mm (orig : 64.32 61.80 -226.08 mm) diff = 0.000 mm
Coordinate transformations established.
Polhemus data for 3 HPI coils added
Device coordinate locations for 3 HPI coils added
Measurement info composed.
Finding samples for /home/ubuntu/mne_data/MNE-brainstorm-data/bst_phantom_ctf/phantom_20uA_20150603_03.ds/phantom_20uA_20150603_03.meg4:
System clock channel is available, checking which samples are valid.
10 x 2400 = 24000 samples from 304 chs
Current compensation grade : 3
Reading 0 ... 23999 = 0.000 ... 10.000 secs...
The sinusoidal signal is generated on channel HDAC006, so we can use that to obtain precise timing.
sinusoid, times = raw[raw.ch_names.index('HDAC006-4408')]
plt.figure()
plt.plot(times[times < 1.], sinusoid.T[times < 1.])
Let’s create some events using this signal by thresholding the sinusoid.
events = np.where(np.diff(sinusoid > 0.5) > 0)[1] + raw.first_samp
events = np.vstack((events, np.zeros_like(events), np.ones_like(events))).T
The CTF software compensation works reasonably well:
raw.plot()
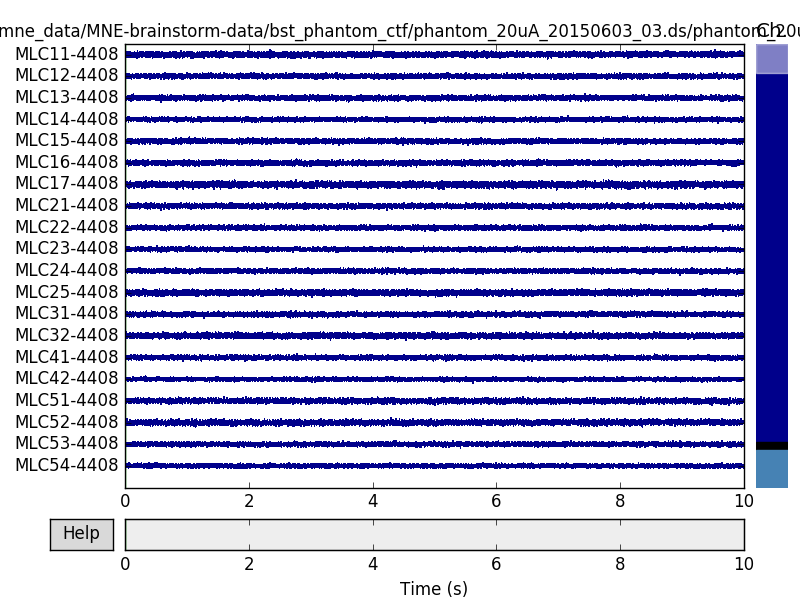
But here we can get slightly better noise suppression, lower localization bias, and a better dipole goodness of fit with spatio-temporal (tSSS) Maxwell filtering:
raw.apply_gradient_compensation(0) # must un-do software compensation first
mf_kwargs = dict(origin=(0., 0., 0.), st_duration=10.)
raw = mne.preprocessing.maxwell_filter(raw, **mf_kwargs)
raw.plot()
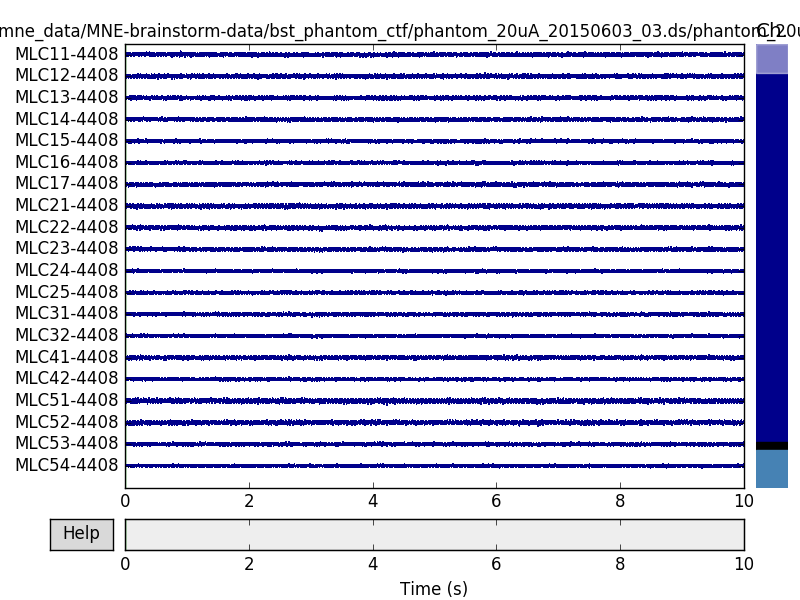
Out:
Compensator constructed to change 3 -> 0
Applying compensator to loaded data
Maxwell filtering raw data
Using loaded raw data
No bad MEG channels
Processing 0 gradiometers and 299 magnetometers
Using origin 0.0, 0.0, 0.0 mm in the head frame
Processing data using tSSS with st_duration=10.0
Computing regularization
Using 86/95 harmonic components for 0.000 (71/80 in, 15/15 out)
Processing 1 data chunks of (at least) 10.0 sec
Projecting 8 intersecting tSSS components for 0.000 - 10.000 sec (#1/1)
[done]
Our choice of tmin and tmax should capture exactly one cycle, so we can make the unusual choice of baselining using the entire epoch when creating our evoked data. We also then crop to a single time point (@t=0) because this is a peak in our signal.
tmin = -0.5 / dip_freq
tmax = -tmin
epochs = mne.Epochs(raw, events, event_id=1, tmin=tmin, tmax=tmax,
baseline=(None, None))
evoked = epochs.average()
evoked.plot()
evoked.crop(0., 0.)
del raw, epochs
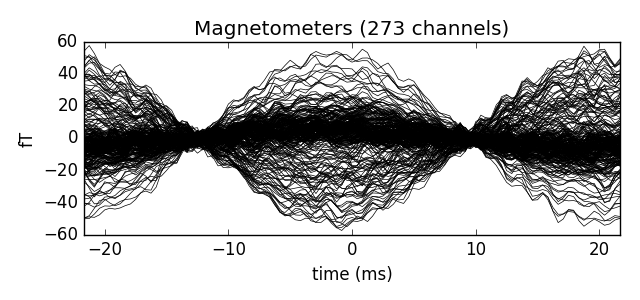
Out:
460 matching events found
0 projection items activated
To do a dipole fit, let’s use the covariance provided by the empty room recording.
raw_erm = read_raw_ctf(erm_path).apply_gradient_compensation(0)
raw_erm = mne.preprocessing.maxwell_filter(raw_erm, coord_frame='meg',
**mf_kwargs)
cov = mne.compute_raw_covariance(raw_erm)
del raw_erm
sphere = mne.make_sphere_model(r0=(0., 0., 0.), head_radius=None)
dip = fit_dipole(evoked, cov, sphere)[0]
Out:
ds directory : /home/ubuntu/mne_data/MNE-brainstorm-data/bst_phantom_ctf/emptyroom_20150709_01.ds
res4 data read.
hc data read.
Separate EEG position data file read.
Quaternion matching (desired vs. transformed):
-0.00 74.50 0.00 mm <-> -0.00 74.50 -0.00 mm (orig : -50.17 57.61 -188.51 mm) diff = 0.000 mm
0.00 -74.50 0.00 mm <-> 0.00 -74.50 0.00 mm (orig : 60.49 -42.15 -190.81 mm) diff = 0.000 mm
74.66 0.00 0.00 mm <-> 74.66 -0.00 0.00 mm (orig : 53.03 61.29 -209.99 mm) diff = 0.000 mm
Coordinate transformations established.
Polhemus data for 3 HPI coils added
Device coordinate locations for 3 HPI coils added
Measurement info composed.
Finding samples for /home/ubuntu/mne_data/MNE-brainstorm-data/bst_phantom_ctf/emptyroom_20150709_01.ds/emptyroom_20150709_01.meg4:
System clock channel is available, checking which samples are valid.
100 x 600 = 60000 samples from 304 chs
Current compensation grade : 3
Compensator constructed to change 3 -> 0
Maxwell filtering raw data
Loading raw data from disk
No bad MEG channels
Processing 0 gradiometers and 299 magnetometers
Using origin 0.0, 0.0, 0.0 mm in the meg frame
Processing data using tSSS with st_duration=10.0
Computing regularization
Using 93/95 harmonic components for 0.000 (78/80 in, 15/15 out)
Processing 10 data chunks of (at least) 10.0 sec
Projecting 8 intersecting tSSS components for 0.000 - 9.998 sec (#1/10)
Projecting 8 intersecting tSSS components for 10.000 - 19.998 sec (#2/10)
Projecting 8 intersecting tSSS components for 20.000 - 29.998 sec (#3/10)
Projecting 8 intersecting tSSS components for 30.000 - 39.998 sec (#4/10)
Projecting 9 intersecting tSSS components for 40.000 - 49.998 sec (#5/10)
Projecting 8 intersecting tSSS components for 50.000 - 59.998 sec (#6/10)
Projecting 8 intersecting tSSS components for 60.000 - 69.998 sec (#7/10)
Projecting 9 intersecting tSSS components for 70.000 - 79.998 sec (#8/10)
Projecting 9 intersecting tSSS components for 80.000 - 89.998 sec (#9/10)
Projecting 8 intersecting tSSS components for 90.000 - 99.998 sec (#10/10)
[done]
Using up to 499 segments
Number of samples used : 59880
[done]
BEM : <ConductorModel | Sphere (no layers): r0=[0.0, 0.0, 0.0] mm>
Sphere model : origin at ( 0.00 0.00 0.00) mm, max_rad = 0.1 mm
Guess grid : 20.0 mm
Guess mindist : 5.0 mm
Guess exclude : 20.0 mm
Using standard MEG coil definitions.
Coordinate transformation: head -> MRI (surface RAS)
1.000000 0.000000 0.000000 0.00 mm
0.000000 1.000000 0.000000 0.00 mm
0.000000 0.000000 1.000000 0.00 mm
0.000000 0.000000 0.000000 1.00
Coordinate transformation: MEG device -> head
0.999939 -0.002039 -0.010868 -2.00 mm
-0.001115 0.959234 -0.282612 -20.74 mm
0.011001 0.282607 0.959173 9.38 mm
0.000000 0.000000 0.000000 1.00
0 bad channels total
Read 273 MEG channels from info
Read 26 MEG compensation channels from info
81 coil definitions read
Coordinate transformation: MEG device -> head
0.999939 -0.002039 -0.010868 -2.00 mm
-0.001115 0.959234 -0.282612 -20.74 mm
0.011001 0.282607 0.959173 9.38 mm
0.000000 0.000000 0.000000 1.00
5 compensation data sets in info
MEG coil definitions created in head coordinates.
Decomposing the sensor noise covariance matrix...
---- Computing the forward solution for the guesses...
Making a spherical guess space with radius 98.6 mm...
Filtering (grid = 20 mm)...
Surface CM = ( 0.0 -0.0 0.0) mm
Surface fits inside a sphere with radius 98.6 mm
Surface extent:
x = -98.6 ... 98.6 mm
y = -98.6 ... 98.6 mm
z = -98.6 ... 98.6 mm
Grid extent:
x = -100.0 ... 100.0 mm
y = -100.0 ... 100.0 mm
z = -100.0 ... 100.0 mm
1331 sources before omitting any.
481 sources after omitting infeasible sources.
Source spaces are in MRI coordinates.
Checking that the sources are inside the bounding surface and at least 5.0 mm away (will take a few...)
4 source space point omitted because of the 5.0-mm distance limit.
Thank you for waiting.
477 sources remaining after excluding the sources outside the surface and less than 5.0 mm inside.
Go through all guess source locations...
[done 477 sources]
---- Fitted : 0.0 ms
1 time points fitted
Compare the actual position with the estimated one.
expected_pos = np.array([18., 0., 49.])
diff = np.sqrt(np.sum((dip.pos[0] * 1000 - expected_pos) ** 2))
print('Actual pos: %s mm' % np.array_str(expected_pos, precision=1))
print('Estimated pos: %s mm' % np.array_str(dip.pos[0] * 1000, precision=1))
print('Difference: %0.1f mm' % diff)
print('Amplitude: %0.1f nAm' % (1e9 * dip.amplitude[0]))
print('GOF: %0.1f %%' % dip.gof[0])
Out:
Actual pos: [ 18. 0. 49.] mm
Estimated pos: [ 18.4 -2.1 44.8] mm
Difference: 4.7 mm
Amplitude: 10.0 nAm
GOF: 96.0 %
Total running time of the script: ( 0 minutes 22.489 seconds)